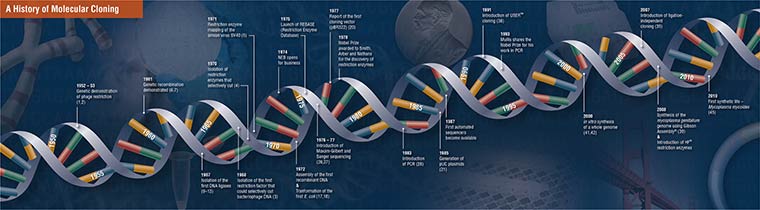
Molecular Cloning
Molecular cloning refers to the process by which recombinant DNA molecules are produced and transformed into a host organism, where they are replicated.
How does it work?
A molecular cloning reaction is typically comprised of the following two components:
- The DNA fragment of interest to be replicated
- A vector/plasmid backbone that contains all of the components for replication in the host
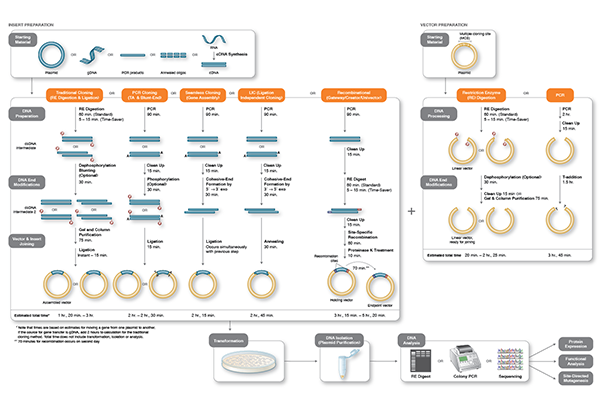
DNA of interest, such as a gene, regulatory element(s), or operon, etc., is prepared for cloning by excising it out of the source DNA using restriction enzymes, copying it using the Polymerase Chain Reaction (PCR), or assembling it from individual oligonucleotides. At the same time, a plasmid vector is prepared in linear form using restriction enzymes or PCR. The plasmid is a small, circular piece of DNA that is replicated within the host, and exists separately from the host’s chromosomal or genomic DNA. By physically joining the DNA of interest to the plasmid vector through phosphodiester bonds, the DNA of interest becomes part of the new recombinant plasmid and is replicated by the host.
Plasmid vectors allow the DNA of interest to be copied in large amounts and, often, provide the necessary control elements to be used to direct transcription and translation of the cloned DNA. As such, they have become the workhorse for many molecular methods, such as protein expression, gene expression studies, and functional analysis of biomolecules.
During the cloning process, the ends of the DNA of interest and the vector have to be modified to make them compatible for joining through the action of a DNA ligase, recombinase, or in vivo DNA repair mechanism. These steps typically utilize enzymes, such as nucleases, phosphatases, kinases and/or ligases. Many cloning methodologies and, more recently, kits have been developed to simplify and standardize these processes.
Synthetic Biology
Synthetic Biology is a more recent expansion of the biotechnology field, in which genes and proteins are viewed as parts or devices, with the goal of re-designing and/or assembling these parts in novel ways to create a new and useful functionality. Recent advances in biofuels generation, production of biochemicals, and understanding the minimal genome all benefit from synthetic biological approaches.
Often these projects rely on the ordered assembly of multiple DNA sequences to create large, artificial DNA structures. To this end, methods have evolved to simplify this process.
NEBuilder® HiFi DNA Assembly and Gibson Assembly® can be used to create many functional DNA structures, from a simple joining of two metabolic genes, all the way up to the creation of an artificial genome.
Further information can be found in our Technical Resources section or at neb.com